Our laboratory is focused on the function of the cerebral cortex which underlies our ability to interact with our environment through sensory perception and voluntary movement. Our research takes a bottom-up approach to understanding how the neural circuits of this massively interconnected network of neurons are functionally organized and how dysfunction in these circuits contributes to neurodegenerative diseases like amyotrophic lateral sclerosis and neuropsychiatric disorders including autism and schizophrenia.
The laboratory is located in the Solomon H. Snyder Department of Neuroscience at Johns Hopkins. The laboratory is also affiliated with the Biochemistry, Cellular and Molecular Biology Graduate Program and the Cellular and Molecular Medicine Program.
Functional organization of the cortex
By combining electrophysiological and optogenetic approaches with anatomical and genetic techniques for identifying cell populations and pathways, we are defining the synaptic interactions among different classes of cortical neurons and determining how long-range and local inputs are integrated within cortical circuits.
Neurodegenerative diseases
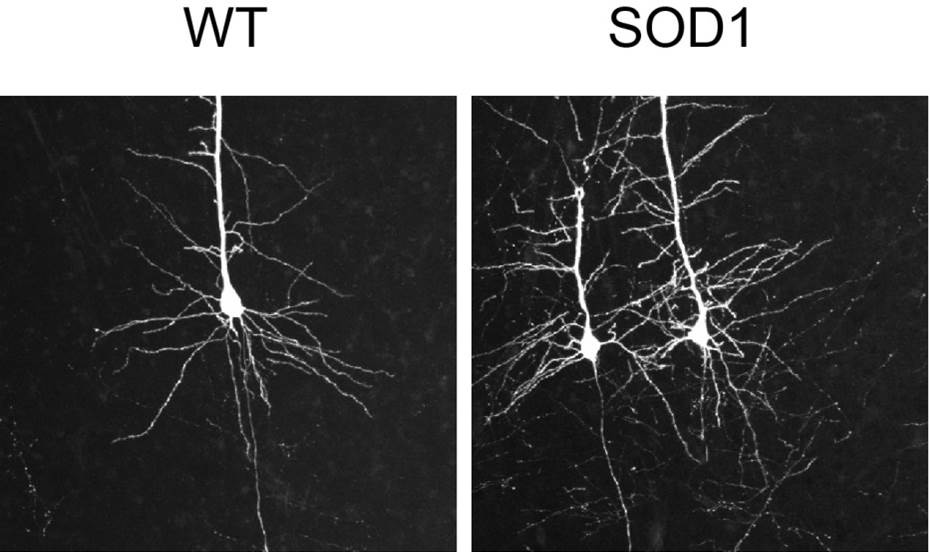
In amyotrophic lateral sclerosis, corticospinal and spinal motor neurons progressively degenerate. We are asking how abnormal activity within cortical circuits contributes to the selective degeneration of corticospinal motor neurons in an effort to identify new mechanisms for treating this disease.
Neuropsychiatric disease
Abnormalities in the organization of cortical circuits and synapses have been identified in genetic and anatomical studies of neuropsychiatric disease. We are interested on the impact these abnormalities have on cortical processing and their contribution to the disordered cognition typical of autism and schizophrenia.
Approaches
We use a variety of techniques to address these questions including:
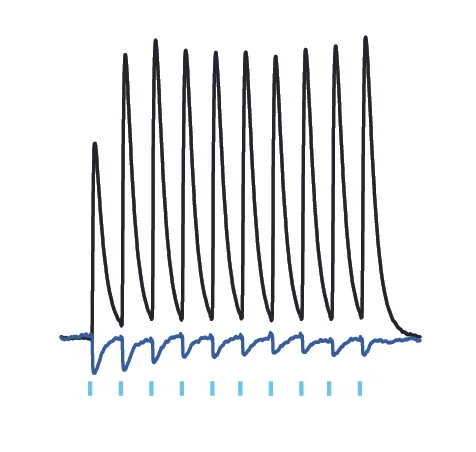
1. Whole-cell patch clamp recording from multiple neurons simultaneously in brain slices to determine the organization of neural circuits and their synaptic properties.
2. Optogenetic approaches to stimulate or inhibit select classes of neurons and interrogate the function of neural circuits.
3. Single cell RNA sequencing.
4. In vivo approaches for monitoring and manipulating neuronal activity.
5. Transgenic mouse lines to control the expression of desired proteins in subsets of neurons.
6. Viral constructs to target protein expression to specific subsets of neurons, allowing us to determine the anatomical relationship among cell types and to probe the functional properties of their connections.